Running compiled Stan models in Shiny
Introduction
The aim of this post is to provide a short step-by-step guide on writing interactive R Shiny-applications that include models written in Stan using rstan and rstantools. The remainder of this post assumes a small amount of working knowledge on writing models in Stan and usage of the package rstan to interface Stan from R.
Demo set-up
For demonstration purposes, let’s start by writing a minimal Stan model file lm.stan:
data {
int<lower=0> N;
vector[N] x;
vector[N] y;
}
parameters {
real alpha;
real beta;
real<lower=0> sigma;
}
model {
y ~ normal(alpha + beta * x, sigma);
}This Stan file encodes a simple linear regression model of the form:
\[ y_i \ \overset{\text{iid}}{\sim} \ N(\alpha + \beta \cdot x_i, \sigma^2), \quad i = 1,\ldots,N \]
Next, we create a small shiny-app contained in a single file app.R (in the same directory as lm.stan) that draws posterior samples from the Stan model in lm.stan with calls to rstan::stan_model() and rstan::sampling():
library(shiny)
library(rstan)
ui <- fluidPage(
sidebarLayout(
sidebarPanel(
numericInput("N", label = "N", value = 10)
),
mainPanel(
plotOutput("posteriors")
)
)
)
server <- function(input, output, session) {
## compile stan model
model <- stan_model(file = "lm.stan")
## draw samples
draws <- reactive({
N <- input$N
sampling(
object = model,
data = list(N = N, x = seq_len(N), y = rnorm(N, seq_len(N), 0.1)),
chains = 2,
iter = 1000
)
})
## plot histograms
output$posteriors <- renderPlot({
req(draws())
op <- par(mfrow = c(1, 2), cex = 1.25)
hist(extract(draws(), "alpha")[[1]], main = bquote("Posterior samples"~alpha), xlab = expression(alpha))
hist(extract(draws(), "beta")[[1]], main = bquote("Posterior samples"~beta), xlab = expression(beta))
par(op)
})
}
shinyApp(ui = ui, server = server)The contents of this shiny-app can be summarized in a simple reactive graph:
New noisy responses \(y_1, \ldots, y_N\) are generated according to \(y_i \overset{\text{iid}}{\sim} N(\alpha + \beta \cdot x_i, \sigma^2)\) with \(x_i = i\) for each \(i = 1,\ldots, N\) and fixed parameters \(\alpha = 0\), \(\beta = 1\) and \(\sigma = 0.1\).
Slow model compilation
A fixed number of 2000 posterior samples for \(\alpha\) and \(\beta\) (and \(\sigma\)) is drawn across two individual chains (i.e. 1000 draws per chain), which should not take more than a few seconds to complete on an ordinary laptop computer, especially if \(N\) is small. However, when launching the shiny-app, it becomes evident that it takes significantly longer to complete drawing any initial posterior samples.
The reason for this is (obviously) that the Stan model has to be recompiled from the lm.stan file whenever we launch the shiny-app in a new R-session due to the call to rstan::stan_model(). Depending on the compiler settings, it takes up to ~1 minute to compile this single Stan model on my laptop computer, which more or less defeats the purpose of creating a shiny-app for interactive use.
Luckily, it is quite simple to avoid this unnecessary computational effort: we just have to compile our Stan models beforehand so that we can sample directly from the already compiled Stan models and skip the compilation step in rstan::stan_model(). Before describing a general R-package approach using rstantools, we start with a simpler approach –suitable for a large set of standard regression models– which is to take advantage of the pre-compiled Stan models in rstanarm.
Pre-compiled models with rstanarm
If the model we wish to sample from is already made available in R via the rstanarm-package, arguably the most straightforward approach to avoid unnecessary Stan model compilation is to use rstanarm’s R wrapper functions to directly sample from a pre-compiled Stan model. Note that if we are fitting a relative standard regression model, there is a decent chance a pre-compiled model version is available in rstanarm. Besides the fact that the Stan models in rstanarm are pre-compiled, the implementations of the Stan programs are likely more robust and computationally stable than any quick Stan program we would implement ourselves.
To sample from a simple linear model as defined in lm.stan with rstanarm, it suffices to remove the call to rstan::stan_model() in app.R and replace rstan::sampling() by a call to rstanarm::stan_lm() or rstanarm::stan_glm()1:
library(shiny)
library(rstanarm)
ui <- fluidPage(
sidebarLayout(
sidebarPanel(
numericInput("N", label = "N", value = 10)
),
mainPanel(
plotOutput("posteriors")
)
)
)
server <- function(input, output, session) {
## draw samples directly
draws <- reactive({
N <- input$N
samples <- stan_glm(
formula = y ~ x,
data = data.frame(x = seq_len(N), y = rnorm(N, seq_len(N), 0.1)),
chains = 2,
iter = 1000
)
as.matrix(samples)[, c(1, 2)]
})
## plot histograms
output$posteriors <- renderPlot({
req(draws())
op <- par(mfrow = c(1, 2), cex = 1.25)
hist(draws()[, 1], main = bquote("Posterior samples"~alpha), xlab = expression(alpha))
hist(draws()[, 2], main = bquote("Posterior samples"~beta), xlab = expression(beta))
par(op)
})
}
shinyApp(ui = ui, server = server)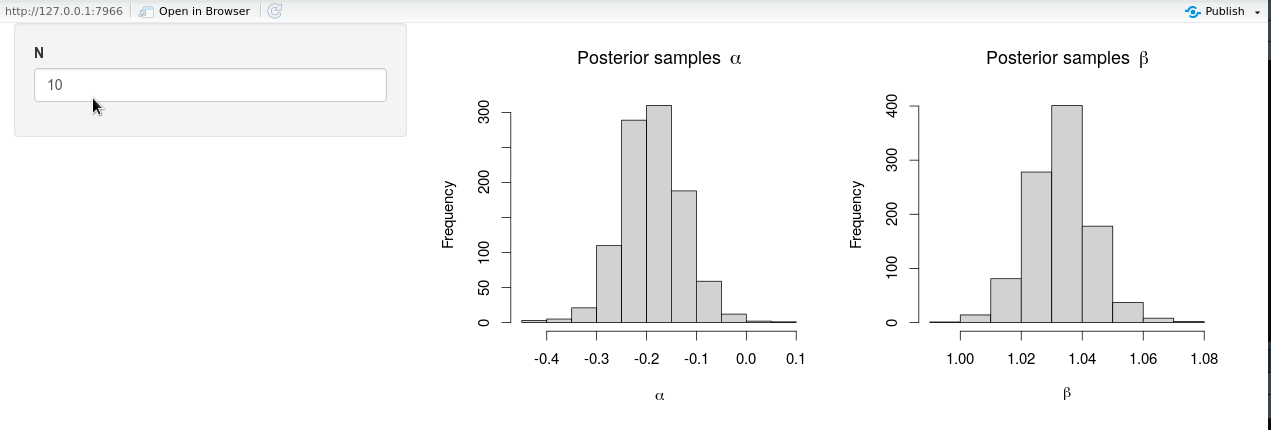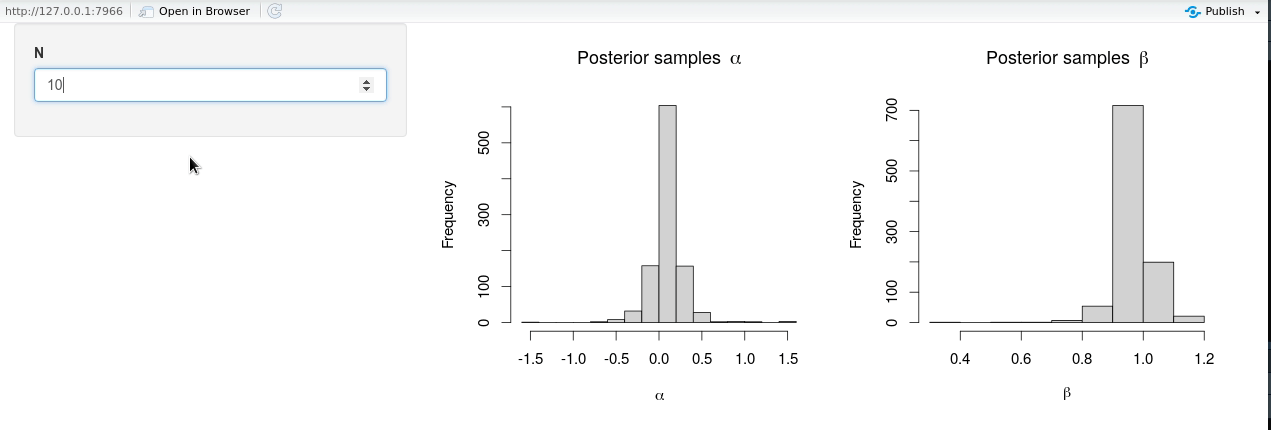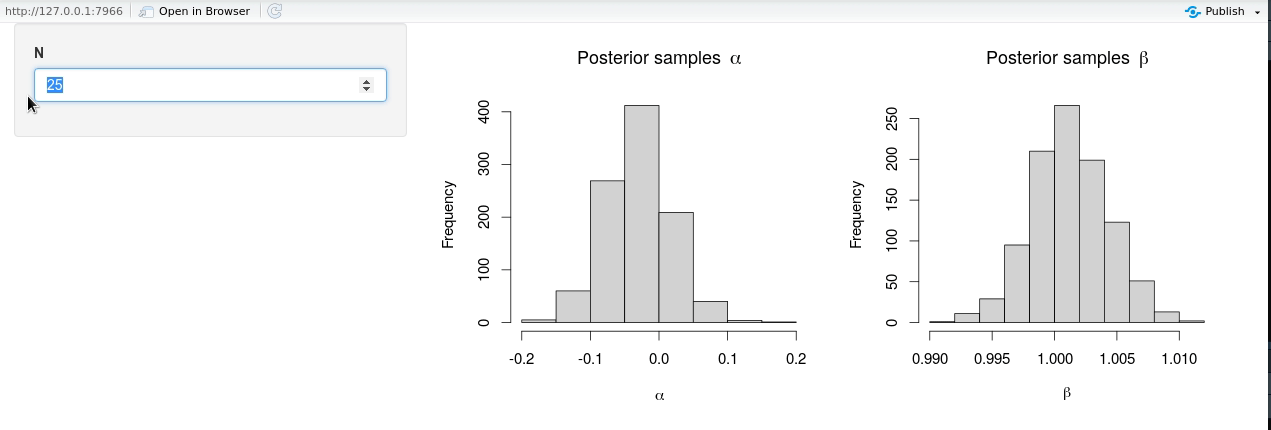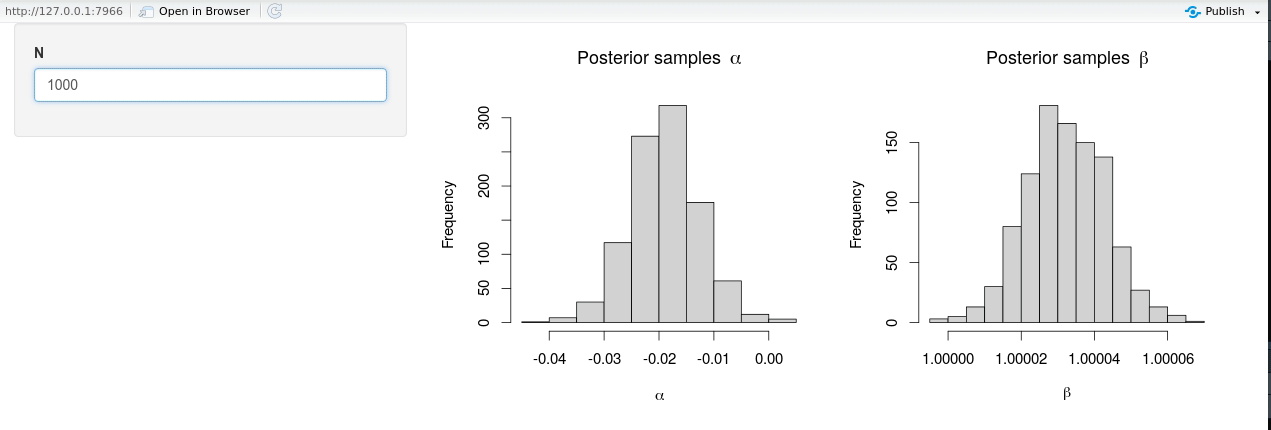
The modified shiny-app no longer exhibits the same lack of responsiveness due to (unnecessary) Stan model recompilation, and it is seen from the animation that new posterior samples are generated almost instantly.
Remark: note that the model used by stan_glm() is not exactly equivalent to the model in lm.stan, since stan_glm() assigns weakly informative priors to the model parameters by default2, whereas (non-informative) uniform priors are used in the original lm.stan file.
That’s great, but what if the model we wish to fit is not available through the rstanarm-package? In this case, we can mimic the same general approach that rstanarm follows: compile the Stan models once on package installation, and directly sample from the pre-compiled models thereafter. As it turns out, we can make use of the excellent rstantools-package for exactly this purpose. The rstantools-package essentially eliminates the effort of setting up a correct R-package structure and only requires us to include the Stan programs that should be compiled with the R-package.
Creating a package with rstantools
To set up a new R-package that should interface Stan, we call rstantools::rstan_create_package() which is roughly similar in use to package.skeleton(), (or usethis::create_package() or Rcpp::Rcpp.package.skeleton() for that matter). The already existing lm.stan file can be included immediately when initializing the package, any new Stan files can be added later to the inst/stan folder. If we do not mind having rstantools as a package dependency, it makes sense to set auto_config = TRUE (the default), which avoids the need to manually reconfigure the package with rstantools::rstan_config() whenever a .stan file is inst/stan are added, removed or modified.
## initialize R-package
rstantools::rstan_create_package(
path = "shinyStanModels",
stan_files = "lm.stan"
)After updating the DESCRIPTION file and roxygenizing the package with roxygen2::roxygenize() or devtools::document(), the R-package can be installed with a call to R CMD INSTALL or devtools::install(). Note that building the package from source takes a while, since this is the moment when the Stan models are compiled and made available to R. The compiled Stan model originating from lm.stan is now directly available in the internal object stanmodels, a named list of S4-objects of class "stanmodel", with each S4-object containing the compiled model version of a single .stan file in the inst/stan folder:
shinyStanModels:::stanmodels[["lm"]]
#> S4 class stanmodel 'lm' coded as follows:
#> data {
#> int<lower=0> N;
#> vector[N] x;
#> vector[N] y;
#> }
#> parameters {
#> real alpha;
#> real beta;
#> real<lower=0> sigma;
#> }
#> model {
#> y ~ normal(alpha + x * beta, sigma);
#> }
class(shinyStanModels:::stanmodels[["lm"]])
#> [1] "stanmodel"
#> attr(,"package")
#> [1] "rstan"At this point we can directly sample from the S4-model objects with rstan::sampling():
system.time({
rstan::sampling(
object = shinyStanModels:::stanmodels[["lm"]],
data = list(N = 10L, x = seq_len(10), y = rnorm(10, seq_len(10), 0.1)),
chains = 2,
iter = 1000
)
})
#>
#> SAMPLING FOR MODEL 'lm' NOW (CHAIN 1).
#> Chain 1:
#> Chain 1: Gradient evaluation took 1.7e-05 seconds
#> Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0.17 seconds.
#> Chain 1: Adjust your expectations accordingly!
#> Chain 1:
#> Chain 1:
#> Chain 1: Iteration: 1 / 1000 [ 0%] (Warmup)
#> Chain 1: Iteration: 100 / 1000 [ 10%] (Warmup)
#> Chain 1: Iteration: 200 / 1000 [ 20%] (Warmup)
#> Chain 1: Iteration: 300 / 1000 [ 30%] (Warmup)
#> Chain 1: Iteration: 400 / 1000 [ 40%] (Warmup)
#> Chain 1: Iteration: 500 / 1000 [ 50%] (Warmup)
#> Chain 1: Iteration: 501 / 1000 [ 50%] (Sampling)
#> Chain 1: Iteration: 600 / 1000 [ 60%] (Sampling)
#> Chain 1: Iteration: 700 / 1000 [ 70%] (Sampling)
#> Chain 1: Iteration: 800 / 1000 [ 80%] (Sampling)
#> Chain 1: Iteration: 900 / 1000 [ 90%] (Sampling)
#> Chain 1: Iteration: 1000 / 1000 [100%] (Sampling)
#> Chain 1:
#> Chain 1: Elapsed Time: 0.027498 seconds (Warm-up)
#> Chain 1: 0.01214 seconds (Sampling)
#> Chain 1: 0.039638 seconds (Total)
#> Chain 1:
#>
#> SAMPLING FOR MODEL 'lm' NOW (CHAIN 2).
#> Chain 2:
#> Chain 2: Gradient evaluation took 4e-06 seconds
#> Chain 2: 1000 transitions using 10 leapfrog steps per transition would take 0.04 seconds.
#> Chain 2: Adjust your expectations accordingly!
#> Chain 2:
#> Chain 2:
#> Chain 2: Iteration: 1 / 1000 [ 0%] (Warmup)
#> Chain 2: Iteration: 100 / 1000 [ 10%] (Warmup)
#> Chain 2: Iteration: 200 / 1000 [ 20%] (Warmup)
#> Chain 2: Iteration: 300 / 1000 [ 30%] (Warmup)
#> Chain 2: Iteration: 400 / 1000 [ 40%] (Warmup)
#> Chain 2: Iteration: 500 / 1000 [ 50%] (Warmup)
#> Chain 2: Iteration: 501 / 1000 [ 50%] (Sampling)
#> Chain 2: Iteration: 600 / 1000 [ 60%] (Sampling)
#> Chain 2: Iteration: 700 / 1000 [ 70%] (Sampling)
#> Chain 2: Iteration: 800 / 1000 [ 80%] (Sampling)
#> Chain 2: Iteration: 900 / 1000 [ 90%] (Sampling)
#> Chain 2: Iteration: 1000 / 1000 [100%] (Sampling)
#> Chain 2:
#> Chain 2: Elapsed Time: 0.017989 seconds (Warm-up)
#> Chain 2: 0.01108 seconds (Sampling)
#> Chain 2: 0.029069 seconds (Total)
#> Chain 2:
#> user system elapsed
#> 0.089 0.000 0.090To keep everything together, we can just as well add the app.R file to the R-package in a folder inst/app. The contents of the app.R file are now similar to our initial attempt, but with the call to rstan::stan_model() left out:
library(shiny)
library(rstan)
ui <- fluidPage(
sidebarLayout(
sidebarPanel(
numericInput("N", label = "N", value = 10)
),
mainPanel(
plotOutput("posteriors")
)
)
)
server <- function(input, output, session) {
## draw samples
draws <- reactive({
N <- input$N
sampling(
object = shinyStanModels:::stanmodels[["lm"]],
data = list(N = N, x = seq_len(N), y = rnorm(N, seq_len(N), 0.1)),
chains = 2,
iter = 1000
)
})
## plot histograms
output$posteriors <- renderPlot({
req(draws())
op <- par(mfrow = c(1, 2), cex = 1.25)
hist(extract(draws(), "alpha")[[1]], main = bquote("Posterior samples"~alpha), xlab = expression(alpha))
hist(extract(draws(), "beta")[[1]], main = bquote("Posterior samples"~beta), xlab = expression(beta))
par(op)
})
}
shinyApp(ui = ui, server = server)The responsiveness of the shiny-app is now the same as in the previous section with the use the rstanarm-package, but we are no longer constrained to only rstanarm’s collection of Stan models.
Models created with brms
Besides rstanarm, the brms-package also provides a flexible interface to build Stan models directly using R syntax. The difference between rstanarm and brms, however, is that brms does not rely on pre-compiled Stan models and compiles generated .stan files on-the-fly. This provides additional flexibility with respect to rstanarm, but also means that calling brms::brm() directly in an interactive shiny-app suffers from the same unresponsiveness as rstan::stan_model().
As a workaround, we can call brms::make_stancode() to return the Stan program generated by brms:
brms::make_stancode(
formula = y ~ x,
data = data.frame(x = numeric(1), y = numeric(1)),
family = "gaussian"
)
#> // generated with brms 2.14.4
#> functions {
#> }
#> data {
#> int<lower=1> N; // total number of observations
#> vector[N] Y; // response variable
#> int<lower=1> K; // number of population-level effects
#> matrix[N, K] X; // population-level design matrix
#> int prior_only; // should the likelihood be ignored?
#> }
#> transformed data {
#> int Kc = K - 1;
#> matrix[N, Kc] Xc; // centered version of X without an intercept
#> vector[Kc] means_X; // column means of X before centering
#> for (i in 2:K) {
#> means_X[i - 1] = mean(X[, i]);
#> Xc[, i - 1] = X[, i] - means_X[i - 1];
#> }
#> }
#> parameters {
#> vector[Kc] b; // population-level effects
#> real Intercept; // temporary intercept for centered predictors
#> real<lower=0> sigma; // residual SD
#> }
#> transformed parameters {
#> }
#> model {
#> // likelihood including all constants
#> if (!prior_only) {
#> target += normal_id_glm_lpdf(Y | Xc, Intercept, b, sigma);
#> }
#> // priors including all constants
#> target += student_t_lpdf(Intercept | 3, 0, 2.5);
#> target += student_t_lpdf(sigma | 3, 0, 2.5)
#> - 1 * student_t_lccdf(0 | 3, 0, 2.5);
#> }
#> generated quantities {
#> // actual population-level intercept
#> real b_Intercept = Intercept - dot_product(means_X, b);
#> }By including this Stan code in the inst/stan folder and rebuilding the R-package, we circumvent the compilation step in brms::brm() and can directly sample from the compiled Stan model with rstan::sampling() as in the previous section. Note that the model input is slightly different, since brms has generated the Stan code for a more general multiple linear model:
system.time({
brms_fit <- rstan::sampling(
object = shinyStanModels:::stanmodels[["brms_lm"]],
data = list(N = 10L, ## number of observations
Y = rnorm(10, seq_len(10), 0.1), ## response vector
K = 2L, ## number of predictors
X = cbind(alpha = rep(1, 10), beta = seq_len(10)), ## predictor matrix
prior_only = FALSE ## set to TRUE to evaluate only the priors
),
chains = 2,
iter = 1000
)
})
#>
#> SAMPLING FOR MODEL 'brms_lm' NOW (CHAIN 1).
#> Chain 1:
#> Chain 1: Gradient evaluation took 6e-06 seconds
#> Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0.06 seconds.
#> Chain 1: Adjust your expectations accordingly!
#> Chain 1:
#> Chain 1:
#> Chain 1: Iteration: 1 / 1000 [ 0%] (Warmup)
#> Chain 1: Iteration: 100 / 1000 [ 10%] (Warmup)
#> Chain 1: Iteration: 200 / 1000 [ 20%] (Warmup)
#> Chain 1: Iteration: 300 / 1000 [ 30%] (Warmup)
#> Chain 1: Iteration: 400 / 1000 [ 40%] (Warmup)
#> Chain 1: Iteration: 500 / 1000 [ 50%] (Warmup)
#> Chain 1: Iteration: 501 / 1000 [ 50%] (Sampling)
#> Chain 1: Iteration: 600 / 1000 [ 60%] (Sampling)
#> Chain 1: Iteration: 700 / 1000 [ 70%] (Sampling)
#> Chain 1: Iteration: 800 / 1000 [ 80%] (Sampling)
#> Chain 1: Iteration: 900 / 1000 [ 90%] (Sampling)
#> Chain 1: Iteration: 1000 / 1000 [100%] (Sampling)
#> Chain 1:
#> Chain 1: Elapsed Time: 0.009366 seconds (Warm-up)
#> Chain 1: 0.006024 seconds (Sampling)
#> Chain 1: 0.01539 seconds (Total)
#> Chain 1:
#>
#> SAMPLING FOR MODEL 'brms_lm' NOW (CHAIN 2).
#> Chain 2:
#> Chain 2: Gradient evaluation took 4e-06 seconds
#> Chain 2: 1000 transitions using 10 leapfrog steps per transition would take 0.04 seconds.
#> Chain 2: Adjust your expectations accordingly!
#> Chain 2:
#> Chain 2:
#> Chain 2: Iteration: 1 / 1000 [ 0%] (Warmup)
#> Chain 2: Iteration: 100 / 1000 [ 10%] (Warmup)
#> Chain 2: Iteration: 200 / 1000 [ 20%] (Warmup)
#> Chain 2: Iteration: 300 / 1000 [ 30%] (Warmup)
#> Chain 2: Iteration: 400 / 1000 [ 40%] (Warmup)
#> Chain 2: Iteration: 500 / 1000 [ 50%] (Warmup)
#> Chain 2: Iteration: 501 / 1000 [ 50%] (Sampling)
#> Chain 2: Iteration: 600 / 1000 [ 60%] (Sampling)
#> Chain 2: Iteration: 700 / 1000 [ 70%] (Sampling)
#> Chain 2: Iteration: 800 / 1000 [ 80%] (Sampling)
#> Chain 2: Iteration: 900 / 1000 [ 90%] (Sampling)
#> Chain 2: Iteration: 1000 / 1000 [100%] (Sampling)
#> Chain 2:
#> Chain 2: Elapsed Time: 0.009642 seconds (Warm-up)
#> Chain 2: 0.007436 seconds (Sampling)
#> Chain 2: 0.017078 seconds (Total)
#> Chain 2:
#> user system elapsed
#> 0.267 0.000 0.267
## alpha is b_Intercept
## beta is b[1]
rstan::summary(brms_fit, pars = c("b_Intercept", "b", "sigma"))[["summary"]]
#> mean se_mean sd 2.5% 25%
#> b_Intercept 0.007348538 0.0055747929 0.13571010 -0.2627963 -0.0720820
#> b[1] 1.001252534 0.0008544422 0.02140303 0.9561450 0.9885503
#> sigma 0.192186382 0.0033915500 0.06206435 0.1123726 0.1499358
#> 50% 75% 97.5% n_eff Rhat
#> b_Intercept 0.005593597 0.08514401 0.3050352 592.6071 1.0014853
#> b[1] 1.001083916 1.01547478 1.0409569 627.4588 0.9997091
#> sigma 0.179320140 0.22264681 0.3182492 334.8790 1.0080279Remark: the Stan code generated by brms contains a bit of unnecessary complexity to sample from a simple linear model, but it does not take any effort to generate this Stan code, as we only need to provide the correct brms model syntax.
A note on deployment
When deploying the shiny-app to e.g. shinyapps.io or RStudio Connect using the rsconnect-package, the R-package generated with rstantools can be made available in a git repository on e.g. github or some other public repository, from which rsconnect is able to fetch and install the R-package when deploying the shiny-app.
Another solution to deploy and host shiny-apps on a server is ShinyProxy, which launches shiny-apps from individual Docker containers. By installing the R-package generated by rstantools when building the Docker image of the shiny-app, we ensure that we can directly sample from our compiled Stan models whenever a new Docker container is started. The following Dockerfile provides a minimal template to install the rstantools-generated R-package from a bundled (.tar.gz) package file and serve the shiny-app at port 3838:
FROM rocker/r-ver:4.0.3
# install system dependencies
RUN apt-get update && \
apt-get install -y --no-install-recommends \
libv8-dev && \
apt-get clean && \
rm -rf /var/lib/apt/lists/
# install R packages (using littler)
# this assumes .tar.gz exists in same folder as Dockerfile
COPY shinyStanModels_0.1.tar.gz ./
RUN install2.r --error shiny rstan rstantools && \
install2.r --error shinyStanModels_0.1.tar.gz && \
rm *.tar.gz
EXPOSE 3838
CMD ["R", "-e", "shiny::runApp(appDir = system.file('app', package = 'shinyStanModels'), port = 3838, host = '0.0.0.0')"]
For simplicity
stan_glm()is used instead ofstan_lm(), asstan_glm()automatically assigns weakly informative priors, whereasstan_lm()expects apriorargument using an additional call toR2().↩︎