Mesmerizing multi-scale Turing patterns in R with Rcpp
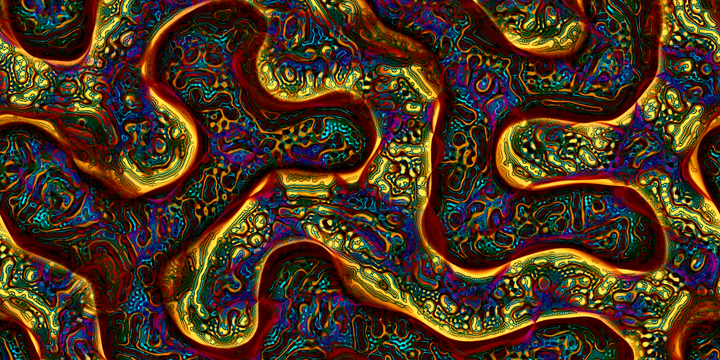
Multi-scale Turing patterns
Turing patterns are a type of reaction-diffusion systems that have attracted much interest as a basic model for the formation of patterns in nature, such as stripes, spots and spirals. The behavior of such diffusion systems was studied by Alan Turing in his classical paper (Turing 1952) and a few decades later re-discovered by (Gierer and Meinhardt 1972), which lead to a more widespread use of the systems in biology as well as other fields1. Turing-like patterns involve activating and inhibiting substances that diffuse through a tissue. If the activator substance diffuses at a lower rate than the inhibitor substance, regular concentration patterns can be formed over time, starting from an almost homogeneous, uniform state. In this post, we follow (McCabe 2010) by focusing on the concentration of a single substance, which serves both as the activator and inhibitor, and diffuses through a discretized surface in the form of a 2D grid of pixels. The change in concentration at a given location (i.e. pixel) is determined by the convolution with respect to a short-range inner 2D kernel (activator) and a long-range outer 2D kernel (inhibitor). If the difference between the activator and inhibitor convolutions is positive, the concentration increases with a certain amount in the next timestep. If the difference is negative, the concentration decreases in the next timestep. (McCabe 2010) extends this basic diffusion model by introducing multiple activator and inhibitor kernels at different scales. The idea is to switch between scales at different locations and timesteps by selecting the scale with the smallest absolute difference between activator and inhibitor convolutions at each location and timestep. The resulting Turing patterns have variation at multiple scales and can produce complex organic looking surfaces.
Implementation
I first came across multi-scale Turing patterns in this post by Jason Rampe, and being quite fascinated by the behavior of the patterns, I thought it would be interesting to write an efficient implementation in R based on 2D convolution with RcppArmadillo (and Rcpp). The R and C++ code is not included with this post, as the current code is not very static and I am adding extra tuning parameters and modifying kernels, colorspaces and interpolation schemes along the way (mostly by trial-and-error) until I am satisfied with the generated results. The generated patterns shown below all use bounded 2D circular or Epanechnikov kernels for the inner (activator) and outer (inhibitor) kernels at up to five different scales, with the radius of the outer kernel around two to three times larger than the inner kernel radius. Kernel convolutions are executed with respect to pre-calculated annuli, where each scale-specific annulus is obtained by taking the difference in densities between the outer and inner kernel. Each evaluated convolution is multiplied by a scale-specific weighting factor and the scale with the smallest absolute variation determines the direction and rate of change in concentration at the next timestep. The main computational bottleneck is the calculation of the kernel convolutions at each location, scale and timestep. These convolutions are performed efficiently in the frequency domain using Armadillo’s forward and backward 2D fast Fourier transforms (arma::fft2 and arma::ifft2). The showcased patterns are evaluated at 1024x1024 pixels per frame across 300 time frames, taking around 1 minute to generate on a modern laptop computer using a single core (Intel i7-8550U CPU, 1.80GHz). The starting frame is initialized by a single non-zero random concentration value at a random location in the grid and the first 50 frames are excluded as warm-up period.
The coloring is inspired by Ricky Reusser’s approach2 in which the scale with the smallest absolute deviation also determines the direction and rate of change in colorspace at the next timestep by moving closer to a fixed color assigned to the selected scale. The patterns below are all generated in the YUV colorspace by mapping the concentration value directly to the luminance component combined with linear or nonlinear 2D interpolation in the U-V color plane. Another colorspace that can be utilized in a similar manner is the YCbCr space by mapping the concentration value to the luma component and considering interpolation in the Cb-Cr color plane.